ESPEI¶
ESPEI, or Extensible Self-optimizing Phase Equilibria Infrastructure, is a tool for automated thermodynamic database development within the CALPHAD method.
The ESPEI package is based on a fork of pycalphad-fitting and uses pycalphad for calculating Gibbs free energies of thermodynamic models. The implementation for ESPEI involves first fitting single-phase data by calculating parameters in thermodynamic models that are linearly described by the single-phase input data. Then Markov Chain Monte Carlo (MCMC) is used to optimize the candidate models from the single-phase fitting to multi-phase zero-phase fraction data. Single-phase and multi-phase fitting methods are described in Chapter 3 of Richard Otis’s thesis.
The benefit of this approach is the automated, simultaneous fitting for many parameters that yields uncertainty quantification, as shown in Otis and Liu High-Throughput Thermodynamic Modeling and Uncertainty Quantification for ICME. Jom 69, (2017).
The name and idea of ESPEI are originally based off of Shang, Wang, and Liu, ESPEI: Extensible, Self-optimizing Phase Equilibrium Infrastructure for Magnesium Alloys Magnes. Technol. 2010 617-622 (2010).
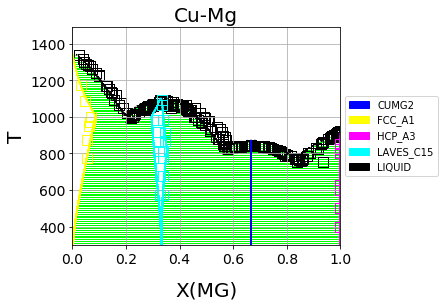
Cu-Mg phase diagram from a database created with and optimized by ESPEI. See the Cu-Mg Example.
Installation¶
Creating a virual environment is highly recommended. ESPEI does require any special compiler, but several dependencies do. Therefore it is suggested to install ESPEI from conda-forge
conda config --add channels conda-forge
conda create -n my_env espei
Alternatively, ESPEI is available from PyPI
pip install espei
or install in develop mode from source
git clone https://github.com/phasesresearchlab/espei.git
cd espei
pip install -e .
Usage¶
ESPEI has two different fitting modes: single-phase and multi-phase fitting. You can run either of these modes or both of them sequentially.
To run either of the modes, you need to have a phase models file that describes the phases in the system using the standard CALPHAD approach within the compound energy formalism.
You also need to describe the data that ESPEI should fit to.
You will need single-phase and multi-phase data for a full run.
Fit settings and all datasets are stored as JSON files and described in detail at the Gathering input data page.
All of your input datasets should be validated by running espei --check-datasets my-input-datasets, where my-input-datasets is a folder of all your JSON files.
The main output result is going to be a database (defaults to out.tdb), an array of the steps in the MCMC chain (defaults to chain.npy), and the an array of the log-probabilities for each iteration and chain (defaults to lnprob.npy).
Single-phase only¶
If you have only heat capacity, entropy and enthalpy data and mixing data (e.g. from first-principles), you may want to see the starting point for your MCMC calculation.
Create an input file called espei-in.yaml.
system:
phase_models: my-phases.json
datasets: my-input-datasets
generate_parameters:
excess_model: linear
ref_state: SGTE91
Then ESPEI can be run by running
espei --input espei-in.yaml
Multi-phase only¶
If you have a database already and just want to do a multi-phase fitting, you can specify a starting TDB file (named my-tdb.tdb) with
system:
phase_models: my-phases.json
datasets: my-input-data
mcmc:
mcmc_steps: 1000
input_db: my-tdb.tdb
The TDB file you input must have all of the degrees of freedom you want as FUNCTIONs with names beginning with VV.
Restart from previous run-phase only¶
If you’ve run an MCMC fitting already in ESPEI and have a chain file called my-previous-chain.npy , then you can resume the calculation with the following input file
system:
phase_models: my-phases.json
datasets: my-input-data
mcmc:
mcmc_steps: 1000
input_db: my-tdb.tdb
restart_chain: my-previous-chain.npy
Full run¶
A minimal full run of ESPEI with single phase fitting and MCMC fitting is done by the following
system:
phase_models: my-phases.json
datasets: my-input-data
generate_parameters:
excess_model: linear
ref_state: SGTE91
mcmc:
mcmc_steps: 1000
Input Customization¶
ESPEI lets you control many aspects of your calculations with the input files shown above. See Writing ESPEI input for a full description of all possible inputs.
FAQ¶
Q: There is an error in my JSON files¶
A: Common mistakes are using single quotes instead of the double quotes required by JSON files. Another common source of errors is misaligned open/closing brackets.
Many mistakes are found with ESPEI’s check-datasets utility.
Run espei check-datasets my-input-datasets on your directory my-input-datasets.
Q: How do I analyze my results?¶
A: By default, ESPEI will create chain.npy and lnprob.npy for the MCMC chain at the end of your run and according to the save interval (defaults to every 20 iterations).
These are created from arrays via numpy.save() and can thus be loaded with numpy.load().
Note that the arrays are preallocated with zeros.
These filenames and settings can be changed using in the input file.
You can then use these chains and corresponding log-probabilities to make corner plots, calculate autocorrelations, find optimal parameters for databases, etc..
Finally, you can use py:mod:espei.plot functions such as multiplot to plot phase diagrams with your input equilibria data and plot_parameters to compare single-phase data (e.g. formation and mixing data) with the properties calculated with your database.
Q: Can I run ESPEI on a supercomputer supporting MPI?¶
A: Yes! ESPEI has MPI support.
To use ESPEI with MPI, you simply call ESPEI in the same way as above with mpirun or whichever MPI software you use.
You also must indicate to ESPEI that it should create an MPI scheduler by setting the input option scheduler: MPIPool in the mcmc heading.
Be aware that mpi4py must be compiled with an MPI-enabled compiler, see the mpi4py installation instructions.
Module Hierarchy¶
fit.pyis the main entry pointparamselect.pyis where all of the fitting happens. This is the core.core_utils.pycontains specialized utilities for ESPEI.utils.pyare utilities with reuse potential outside of ESPEI.plot.pyholds plotting functions
License¶
ESPEI is MIT licensed. See LICENSE.
Tutorials
Reference